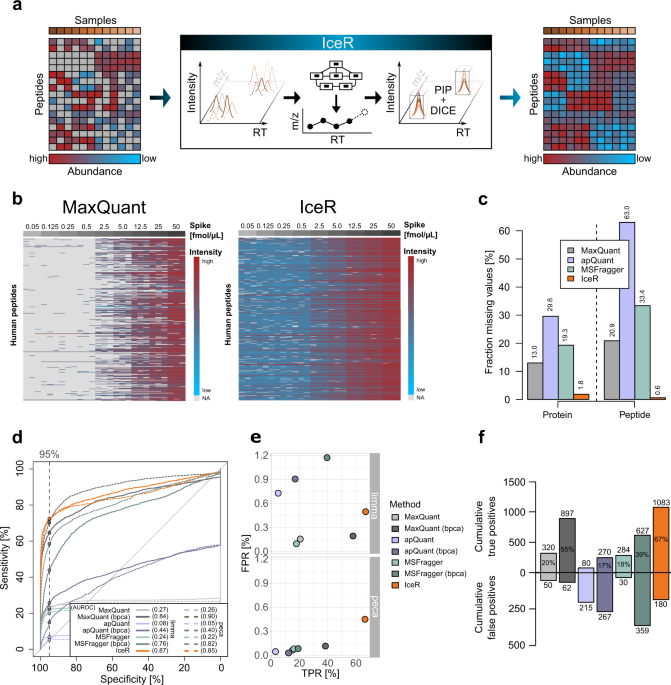
IceR improves proteome coverage and data completeness in global and single-cell proteomics | Nature Communications

Hybrid-DIA: intelligent data acquisition integrates targeted and discovery proteomics to analyze phospho-signaling in single spheroids | Nature Communications
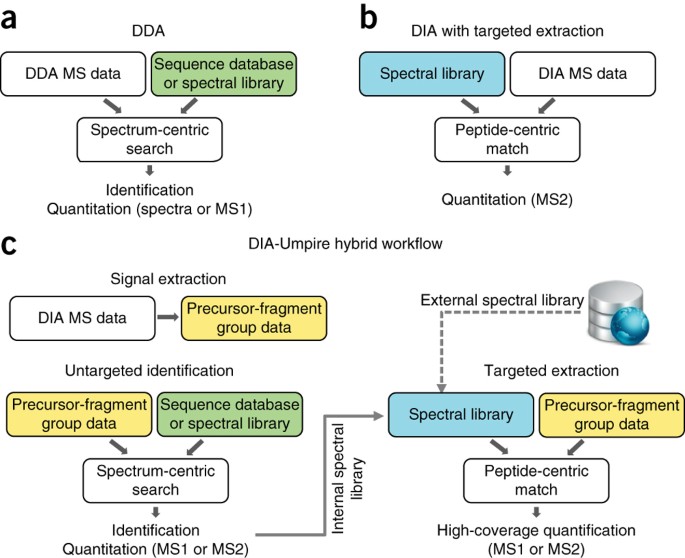
DIA-Umpire: comprehensive computational framework for data-independent acquisition proteomics | Nature Methods

Implementing the reuse of public DIA proteomics datasets: from the PRIDE database to Expression Atlas | Scientific Data
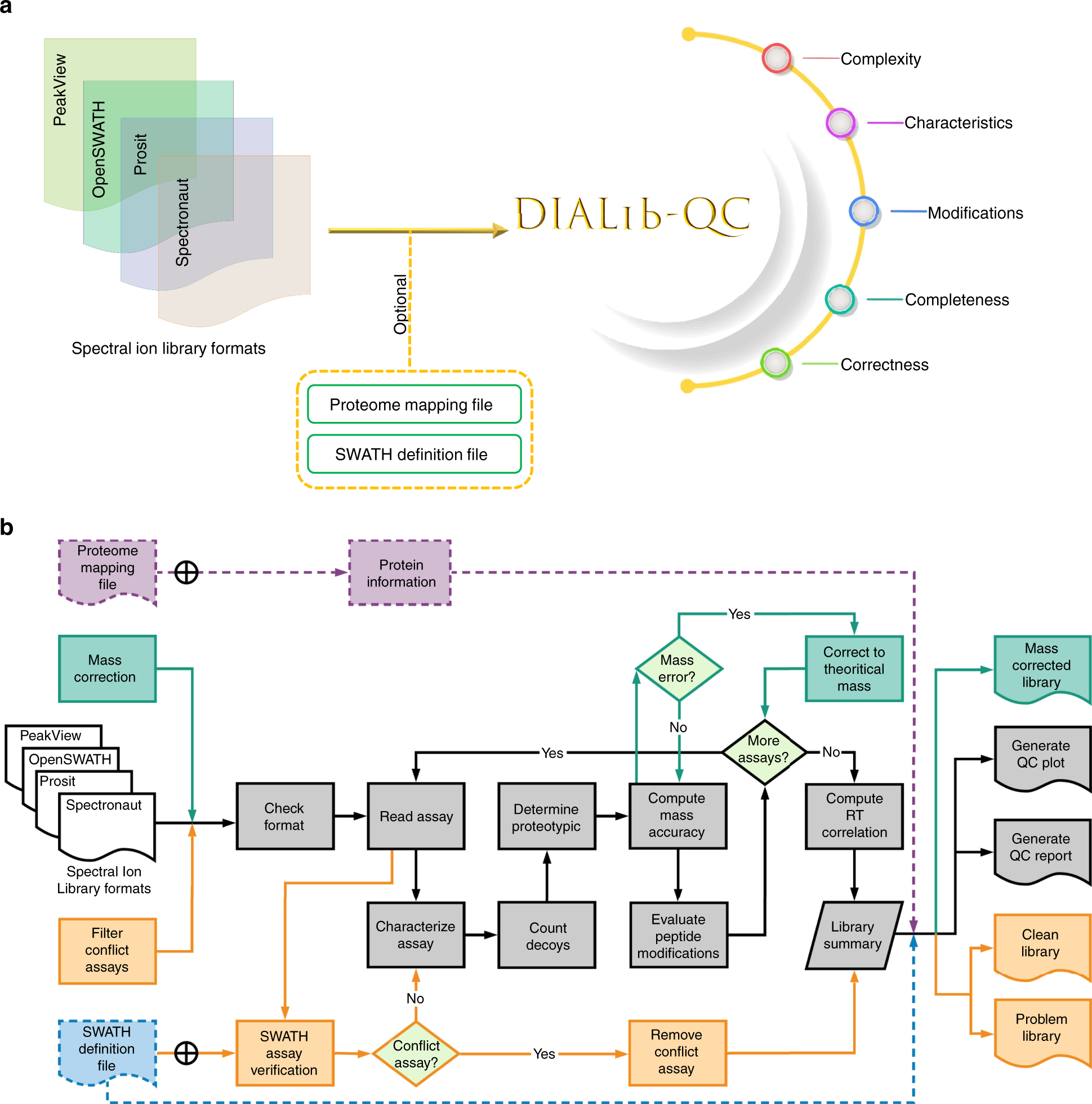
DIALib-QC an assessment tool for spectral libraries in data-independent acquisition proteomics | Nature Communications
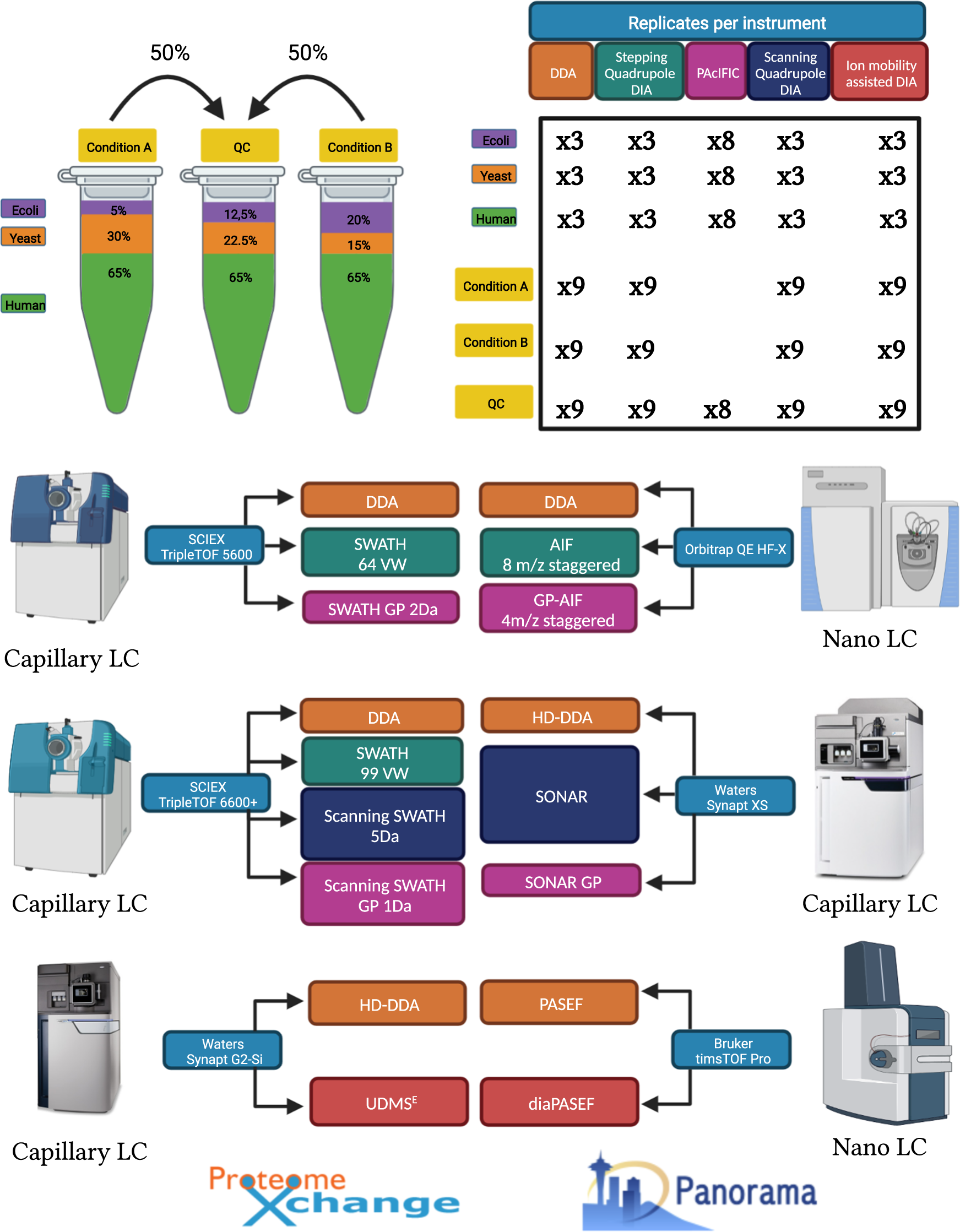
A comprehensive LFQ benchmark dataset on modern day acquisition strategies in proteomics | Scientific Data
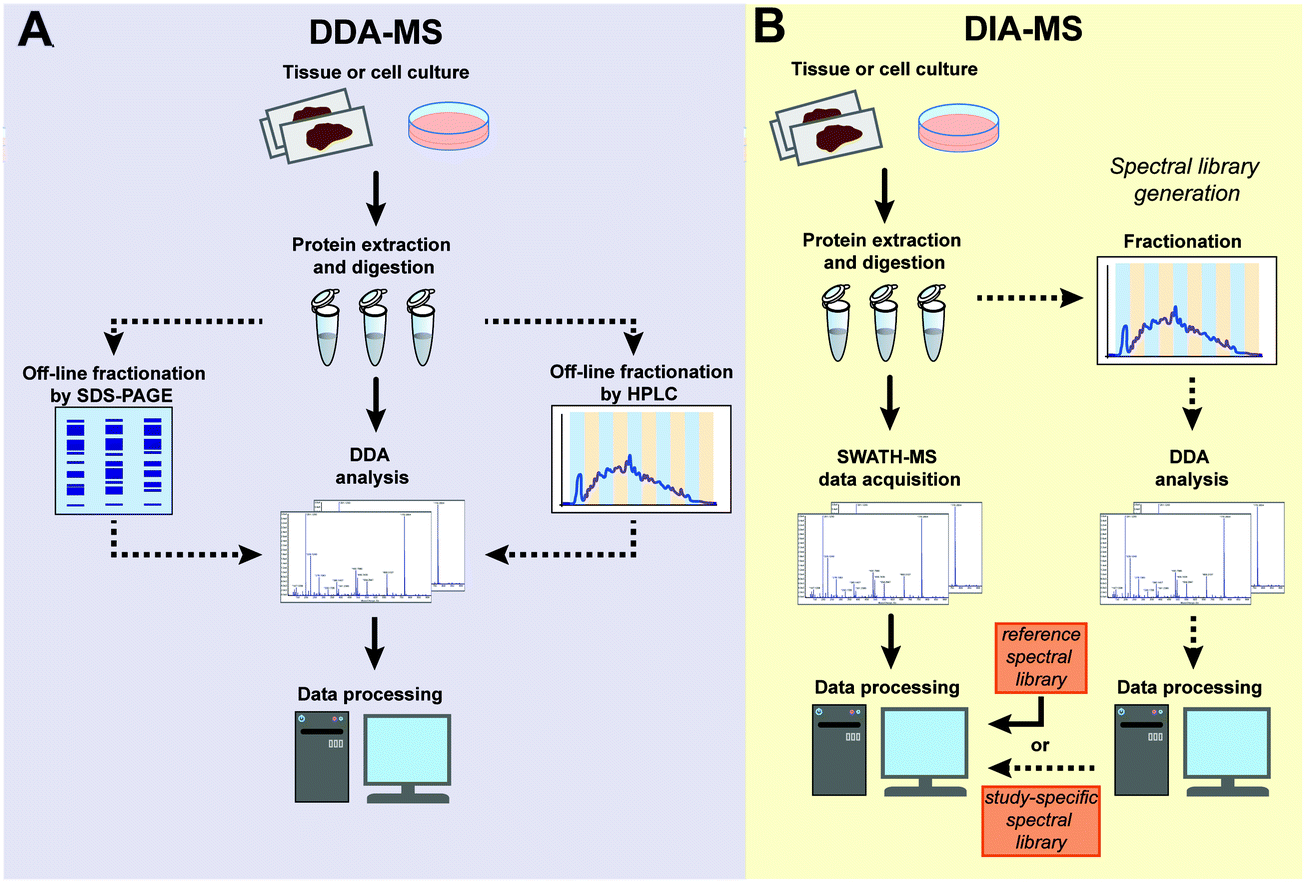
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H
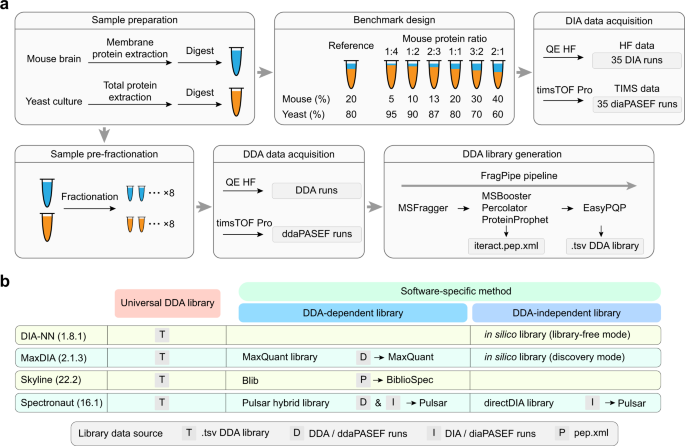
Benchmarking commonly used software suites and analysis workflows for DIA proteomics and phosphoproteomics | Nature Communications
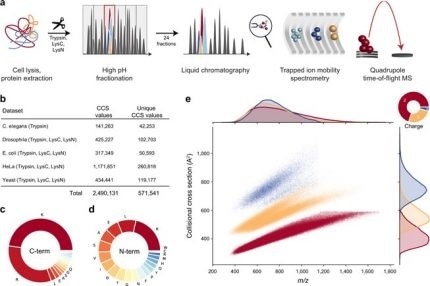
New Nature Communications publication by Mann & Theis Groups harnesses the benefits of large-scale peptide

Use of Hybrid Data-Dependent and -Independent Acquisition Spectral Libraries Empowers Dual-Proteome Profiling | Journal of Proteome Research
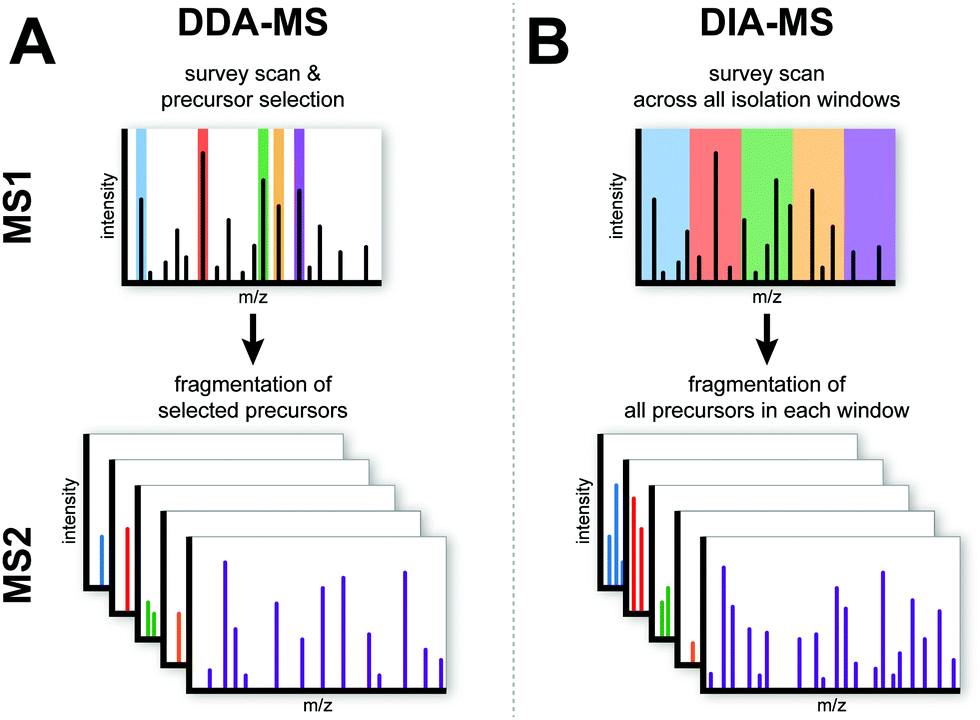
Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology - Molecular Omics (RSC Publishing) DOI:10.1039/D0MO00072H

Chromatogram libraries improve peptide detection and quantification by data independent acquisition mass spectrometry | Nature Communications
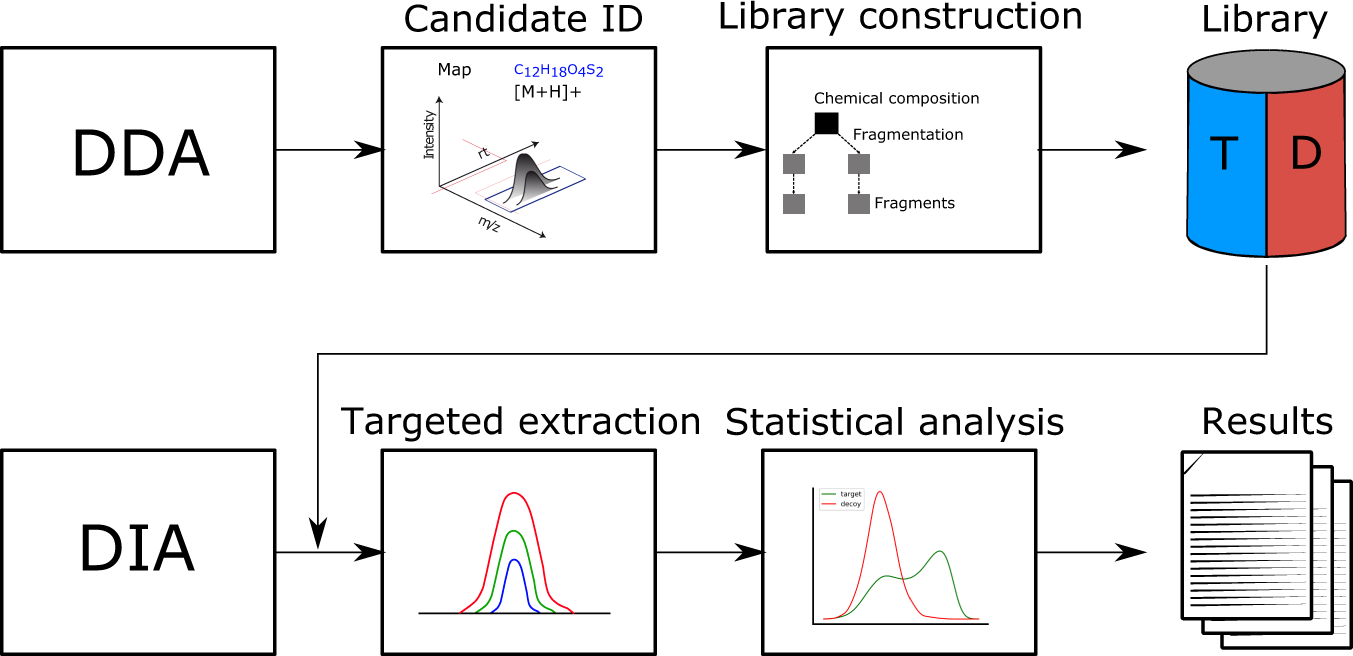
DIAMetAlyzer allows automated false-discovery rate-controlled analysis for data-independent acquisition in metabolomics | Nature Communications